Vahaduo is one of the most powerful online tools to explore your deep ancestry using G25 coordinates, a set of principal components developed by Davidski from the Eurogenes project. In this tutorial, we’ll walk you through how to use Vahaduo to model your ancestry in just a few simple steps.
Levant_South,0.085367,0.148674,-0.057398,-0.092508,-0.009602,-0.035029,-0.002209,-0.008169,0.016321,0.008237,0.009159,-0.009921,0.020485,0.011863,-0.005537,0.002121,-0.011239,0.001014,0.002112,-0.003652,-0.000524,0.003116,0.000567,-0.00294,0.004646
Levant_North,0.088149,0.145785,-0.048858,-0.078848,-0.012891,-0.03074,-0.000496,-0.004282,0.005181,0.010225,0.006658,-0.004529,0.01039,0,-0.010315,0.008471,-0.002101,0.001788,0.002542,-0.000125,0.000763,-0.000302,-0.00141,0.002356,-0.001211
Jordan_Plateau,0.048944,0.131206,-0.047442,-0.07442,-0.012618,-0.027276,-0.004935,-0.003231,0.014889,0.000838,0.006463,-0.008722,0.015996,-0.000936,-0.001113,0.009812,0.000104,0.000887,-0.002011,0.004227,0.001622,0.002893,-0.001134,0.002506,-0.002443
Syria_Interior,0.063236,0.127167,-0.048649,-0.062806,-0.01773,-0.020127,-0.003486,-0.002269,-0.000568,-0.001053,0.002806,-0.005603,0.011422,0.000986,-0.002217,0.004773,-0.004665,-0.000739,0.00192,0.001438,0.001608,0.000927,-0.002232,-0.000522,-0.000246
Kurdistan_Highlands,0.092295,0.112234,-0.062164,-0.038063,-0.037469,-0.004055,0.004428,-0.005603,-0.025112,-0.012856,0.002509,-0.000688,0.005105,0.000122,0.001582,0.010425,-0.005015,0.001254,0.004117,-0.009066,-0.000427,-0.001068,-0.00016,-0.001486,0.004059
Italy_North,0.12384,0.150705,0.030698,-0.011757,0.036191,-0.002677,0.001081,0.001708,0.010758,0.027663,-0.001397,0.006804,-0.012844,-0.006276,-0.003474,-0.004163,-0.000365,0.000659,0.005154,-0.005553,-0.001048,-0.000025,-0.00419,0.003784,0.000958
Italy_South,0.101682,0.146066,-0.003834,-0.038330,0.018618,-0.012597,-0.001293,0.000307,0.010464,0.021808,0.005359,0.003497,-0.004188,0.002500,-0.004094,-0.004840,-0.002368,-0.000591,0.001801,-0.005003,0.000395,0.001752,-0.002547,-0.000603,0.000319
Italy_Central,0.118810,0.147784,0.013882,-0.021011,0.025294,-0.008938,-0.001063,-0.001626,0.005649,0.023239,-0.000309,0.006066,-0.011475,-0.003703,-0.002243,-0.000814,0.002154,0.000290,0.003334,-0.003829,-0.001967,0.001407,-0.000734,0.003196,-0.001226
Arabian_Gulf,0.053393,0.141712,-0.064899,-0.114607,-0.012422,-0.046930,-0.012242,-0.008727,0.050424,-0.005169,0.017405,-0.032426,0.063830,0.003140,0.003924,0.029037,-0.022568,0.004814,-0.001394,0.031481,0.013023,0.018154,-0.007462,0.008884,-0.010113
Maghreb_East,-0.048517,0.131384,-0.015085,-0.075138,0.019619,-0.030678,-0.022091,0.005019,0.054378,0.019795,0.005440,-0.008037,0.026740,-0.010184,0.013046,-0.007143,-0.003716,-0.012922,-0.029005,0.005174,-0.010450,-0.022829,0.018117,-0.003344,0.000210
Maghreb_West,-0.085367,0.130213,-0.012822,-0.070881,0.021132,-0.032940,-0.027783,0.009769,0.055403,0.024399,0.009509,-0.009025,0.026990,-0.014466,0.016302,-0.006188,-0.002767,-0.017033,-0.038422,0.009435,-0.009844,-0.029512,0.020911,-0.001125,0.005921
India_Bengal_Delta,0.041713,-0.119503,-0.145613,0.099722,-0.052951,0.061758,-0.001611,0.011769,0.041885,0.025695,-0.004585,0.001389,-0.000044,0.000069,-0.006151,-0.006579,0.004556,-0.000693,-0.002910,0.004521,0.000907,0.002811,0.001316,0.004852,0.000257
Italy_Sardinia,0.121687,0.167285,0.028490,-0.050652,0.060151,-0.022134,-0.003952,0.002496,0.041574,0.077351,-0.000059,0.016649,-0.028664,-0.012974,-0.013572,-0.003001,0.011403,-0.001347,0.001680,-0.012995,-0.002121,-0.001102,-0.010084,-0.021383,0.000337
Iberia_Basque,0.126424,0.148739,0.055679,0.008790,0.055450,0.000249,-0.001754,0.000025,0.029539,0.042923,-0.005214,0.010801,-0.024986,-0.019493,0.015807,0.002614,-0.005630,0.003172,-0.002375,-0.001791,0.008819,0.002204,-0.006479,-0.008654,0.001150
Scotland_Highlands,0.131466,0.134232,0.062265,0.047043,0.039106,0.017281,0.003550,0.005176,0.003572,0.002825,-0.005614,0.005047,-0.011680,-0.011973,0.023024,0.003125,-0.011255,0.002774,0.002550,0.001452,0.004225,0.003308,-0.000656,0.013556,-0.000770
Ireland_Midlands,0.133361,0.134098,0.061169,0.048864,0.037788,0.019352,0.003293,0.004713,0.003568,0.002927,-0.006971,0.005840,-0.014189,-0.014076,0.025935,0.005215,-0.011145,0.001894,0.000597,0.001776,0.005114,0.001279,0.000293,0.014407,0.000661
Fennoscandia_Saami_North,0.111262,-0.030212,0.111722,0.079781,-0.009329,0.008297,0.008137,0.013860,0.002966,-0.032096,0.022288,-0.007475,0.018071,-0.019826,-0.004004,0.002693,0.001157,-0.001489,-0.005986,0.004494,0.015816,0.001345,-0.002565,0.002658,0.000344
Baltic_Region,0.134197,0.118309,0.088265,0.086387,0.042993,0.031892,0.011986,0.014261,-0.000879,-0.032666,-0.001754,-0.013084,0.021340,0.026369,-0.008612,0.002883,0.001467,-0.001432,0.001477,0.003258,-0.000705,-0.003790,0.006667,-0.004290,0.002844
Slavic_West,0.131500,0.130566,0.065176,0.052498,0.039667,0.020318,0.007511,0.009763,0.001870,-0.013720,-0.003017,-0.005158,0.009571,0.015200,-0.002641,0.000220,0.000452,-0.000110,0.003131,0.002092,-0.002198,-0.001640,0.006059,0.000299,0.000753
Slavic_East,0.132262,0.123623,0.075022,0.069596,0.040233,0.026569,0.010732,0.013322,-0.000273,-0.024602,-0.002696,-0.009212,0.019326,0.027231,-0.011265,-0.003801,-0.000965,0.000372,0.004140,-0.001051,-0.004234,-0.004608,0.007724,-0.006860,0.000695
Slavic_South,0.126878,0.136504,0.038596,0.016325,0.030209,0.006395,0.005480,0.005030,-0.000013,0.000819,-0.001097,-0.001143,0.003581,0.012310,-0.012595,-0.001277,0.006066,0.000329,0.003998,-0.001652,-0.006851,-0.001614,0.004753,-0.000709,-0.000888
Scandinavia_Main,0.131844,0.130135,0.069024,0.055314,0.040630,0.020039,0.005233,0.007596,0.004412,-0.003836,-0.004735,0.002763,-0.006929,-0.006865,0.017838,0.006180,-0.006275,0.002326,0.003069,0.004490,0.006031,0.003017,0.000944,0.013158,0.000115
Central_Asian,0.079169,-0.074084,-0.004858,0.001093,-0.038165,0.000947,0.007008,0.004693,-0.016117,-0.010328,-0.013142,-0.002249,0.001782,-0.003628,0.004269,0.004946,-0.001990,-0.000068,0.000811,-0.001974,-0.007839,-0.002260,-0.003832,-0.000914,0.001899
Africa_East,-0.296623,0.093429,-0.026851,-0.069736,0.000954,-0.034080,-0.020493,0.005769,0.115392,-0.079674,-0.008948,-0.006669,0.005218,-0.001665,0.024728,-0.018616,0.014864,-0.001976,0.009980,-0.003839,0.000599,0.004946,-0.002736,-0.001241,-0.002790
Africa_West,-0.618275,0.064848,0.020924,0.014087,0.001303,0.009026,-0.040128,0.043268,-0.040308,0.028458,0.005017,-0.001429,0.019643,-0.000017,0.011769,-0.012471,0.008384,-0.000367,0.001953,-0.003110,-0.000044,0.000251,0.001318,-0.000286,0.000476
China_South,0.018970,-0.448694,-0.026231,-0.065103,0.102497,0.047659,0.000685,-0.005468,-0.016731,-0.007887,-0.020989,-0.004601,0.003667,-0.002760,-0.001195,0.002342,0.001239,-0.001481,-0.004165,-0.011669,0.013653,0.009438,0.014930,-0.000288,0.003906
China_North,0.025285,-0.446253,0.010344,-0.063747,0.047152,0.020917,0.006026,0.003115,-0.012344,0.003046,-0.071346,-0.007868,0.007922,-0.007235,-0.006873,0.000464,-0.002412,0.000136,-0.003187,-0.006065,0.010580,0.003604,0.010062,0.000361,-0.002164
Japan_Korea,0.022788,-0.450483,0.015616,-0.061619,0.037520,0.010473,0.004311,0.002038,-0.008017,0.008647,-0.074547,-0.008060,0.010837,-0.006437,-0.010035,-0.003481,0.000597,0.002734,0.000754,-0.008281,0.021635,-0.010477,0.006157,0.002184,-0.029733
Aegean,0.106977,0.145820,-0.019480,-0.050973,0.004481,-0.016813,0.003041,-0.003036,-0.002691,0.014768,0.003389,0.001825,-0.003125,0.001705,-0.011953,-0.001788,0.003985,0.002985,0.004857,-0.004822,-0.004379,-0.000032,0.000495,0.000287,-0.002553
Caucasus_North,0.110689,0.100057,-0.032791,-0.021497,-0.035116,-0.000369,0.009279,-0.004390,-0.054759,-0.025349,-0.001210,0.007732,-0.019783,0.001725,0.004796,-0.015229,0.008029,-0.005061,-0.009802,0.017975,0.008991,0.001495,0.002149,-0.004110,-0.003012
Caucasus_South,0.105774,0.134032,-0.055219,-0.054578,-0.032641,-0.012760,0.006014,-0.005239,-0.040846,-0.010136,0.002464,0.005701,-0.010135,0.003165,-0.001831,-0.008953,0.003382,-0.001319,-0.002189,0.003696,0.004721,0.002156,-0.000175,-0.003806,-0.000038
Iran_Highlands,0.089920,0.101959,-0.078969,-0.021189,-0.058749,0.005912,0.007614,-0.005054,-0.041375,-0.021267,-0.001364,-0.001888,-0.000074,0.001156,0.005592,0.010713,-0.006376,0.003281,0.004198,-0.014019,-0.000911,-0.007691,-0.001553,-0.005374,0.005041
Vietnam_Cambodia,0.013924,-0.394296,-0.071055,-0.030270,0.121971,0.063192,-0.003231,-0.008204,-0.004650,-0.012717,0.054398,0.004311,-0.004274,0.001239,0.001375,-0.003799,-0.000799,-0.003068,-0.004033,0.007158,-0.008749,0.011337,-0.001939,0.002655,0.027786
Balkan_General,0.126371,0.137956,0.037429,0.012397,0.030784,0.005491,0.004819,0.004905,0.000309,0.002821,-0.000739,-0.000617,0.001478,0.011618,-0.013483,-0.000552,0.007324,-0.000194,0.005223,-0.002003,-0.007094,-0.000814,0.004881,0.000656,-0.000934
Iran_Central,0.087661,0.094267,-0.069210,-0.017737,-0.048842,0.003918,0.004946,-0.003495,-0.030018,-0.017584,-0.000123,-0.001944,0.003852,-0.003345,0.006556,0.014096,-0.003763,0.002059,0.002284,-0.010313,-0.000816,-0.004195,-0.000479,-0.004484,0.005108
Iraq_Mesopotamia,0.059507,0.117923,-0.062546,-0.057779,-0.028827,-0.015882,-0.002720,-0.004616,-0.001845,-0.006626,0.005535,-0.006638,0.015714,0.000690,0.001925,0.009210,-0.009024,0.003238,0.002785,-0.001127,0.000481,0.000388,-0.001176,0.000377,0.000824
Anatolia_Central,0.106588,0.125382,-0.012526,-0.020130,-0.008925,0.000757,0.004347,-0.000404,-0.014901,-0.004764,-0.000934,0.000626,-0.003106,0.004055,-0.002734,-0.000999,-0.005109,-0.001294,0.003484,-0.004596,-0.002197,0.003670,0.004296,-0.001188,0.003071
Horn_Africa,-0.259101,0.100063,-0.031103,-0.077263,0.001212,-0.036875,-0.014521,-0.000175,0.106946,-0.069999,-0.004610,-0.010118,0.012227,0.000066,0.022763,-0.014639,0.012763,0.001056,0.007802,0.000304,0.002361,0.005730,-0.003537,0.001916,-0.003129
South_Asia_Interior,0.062004,-0.033422,-0.122184,0.088524,-0.069736,0.054054,0.001392,0.007429,0.014921,0.007249,-0.006843,-0.000216,-0.000681,-0.003418,0.004907,0.003220,-0.002445,-0.000158,0.000440,-0.004797,0.000057,-0.002572,0.001733,-0.000245,-0.000003
Tibetan_Himalayas,0.028024,-0.342713,-0.024163,-0.009564,0.003836,0.018216,0.005478,0.006533,0.007485,0.012508,-0.064812,-0.007506,0.008052,-0.002405,-0.007222,-0.004168,0.000450,-0.001811,-0.006811,0.002671,0.000668,0.018559,0.005071,0.000696,0.025524
Southeast_Asia,0.012830,-0.352039,-0.091323,-0.003500,0.092988,0.058393,-0.003671,-0.005084,0.007170,-0.002043,0.047778,0.004667,-0.005592,0.004102,0.000052,-0.002126,0.000456,-0.000973,-0.000003,0.007962,-0.004814,0.004841,-0.006108,0.000392,0.009558
Native_America,0.054741,-0.302734,0.112213,0.088447,-0.105646,-0.016245,-0.264462,-0.316927,-0.009644,-0.016155,-0.001006,-0.002249,0.001469,0.017441,-0.009351,0.003670,0.005187,0.001061,0.002602,0.001071,-0.001045,0.002976,0.001001,0.002281,0.000333
Oceania_Island,-0.019024,-0.216536,-0.189635,0.220962,0.174225,-0.359067,-0.001856,0.003571,-0.034284,-0.013469,-0.007158,0.001304,0.000277,-0.004577,0.004112,0.000000,-0.003888,0.000693,0.001421,-0.004285,0.003305,-0.001179,-0.000196,-0.000428,-0.006271
Iberia_Main,0.126690,0.150851,0.035253,0.005179,0.048573,-0.002410,-0.001640,0.000799,0.027414,0.039412,-0.003037,0.010125,-0.021799,-0.017713,0.013424,0.002840,-0.005301,0.002990,-0.000560,-0.001404,0.007108,0.001349,-0.005866,-0.006632,0.001175
France_South,0.123777,0.152449,0.030754,0.000456,0.045128,-0.001755,-0.001038,0.000975,0.024667,0.036681,-0.002915,0.009838,-0.020888,-0.016918,0.012065,0.002737,-0.004918,0.002767,-0.000797,-0.001692,0.006479,0.001243,-0.005505,-0.006097,0.001113
France_North,0.130248,0.144172,0.046672,0.025342,0.042005,0.009144,0.003632,0.004508,0.012086,0.013814,-0.004601,0.004116,-0.010319,-0.011225,0.018950,0.003943,-0.009148,0.002140,0.002150,0.001140,0.004174,0.002532,-0.001645,0.008721,-0.000061
Germany_Main,0.130376,0.135369,0.054563,0.037280,0.041414,0.014738,0.005178,0.006681,0.006947,0.006015,-0.005271,0.002440,-0.007557,-0.008601,0.020931,0.004099,-0.008752,0.002073,0.002978,0.002219,0.004787,0.002600,-0.001017,0.010299,-0.000236
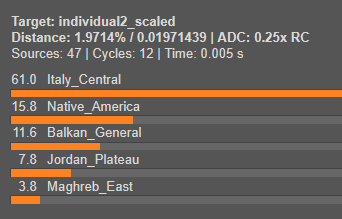
This situation is common and does not invalidate the model, it simply suggests refining the reference list or exploring deeper ancestry layers.
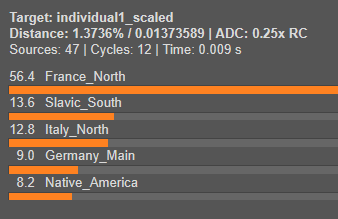
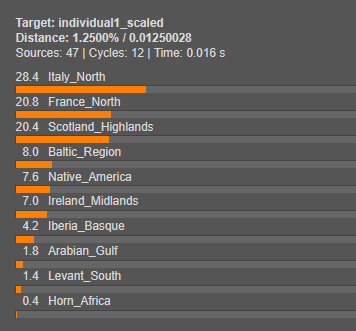
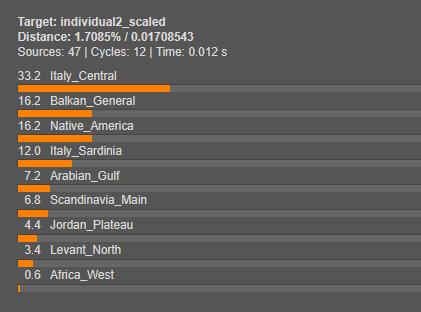
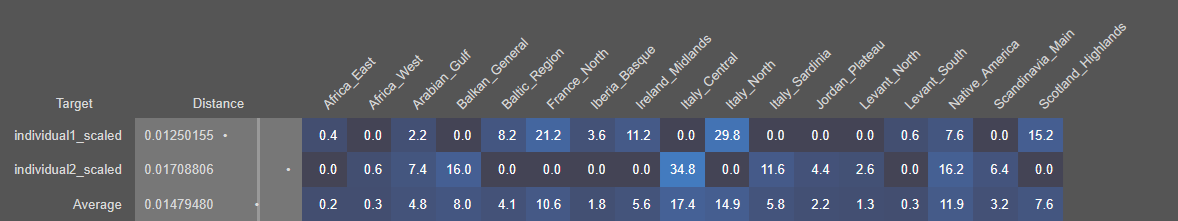
We will now model both Individual 1 and Individual 2 using Single Mode. This step will reveal what their genetic makeup is composed of by estimating the proportions contributed by each source population.
Given all the results obtained through Distance Mode, Single Mode, and Multi Mode, we can draw the following conclusions:
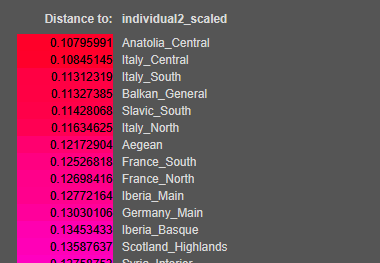
These distinctions illustrate how tools like Vahaduo G25 can uncover both shared ancestry and nuanced individual differences, making them valuable for personal exploration and population studies.
Vahaduo is a simple yet powerful tool to visualize your ancestry using G25 coordinates. By testing different ancient and modern populations, you can build detailed models of your genetic past.